Homogeneous assays utilizing Fluorogen Activating Peptide (FAP) technology have proven to be very valuable for measuring GPCR internalization and trafficking.
You can download our full list of available cell lines expressing FAP-tagged GPCRs here: Available GPCR Cell Lines
Advanced GPCR internalization assays can be developed by utilizing two different fluorogens that are activated by the same FAP.
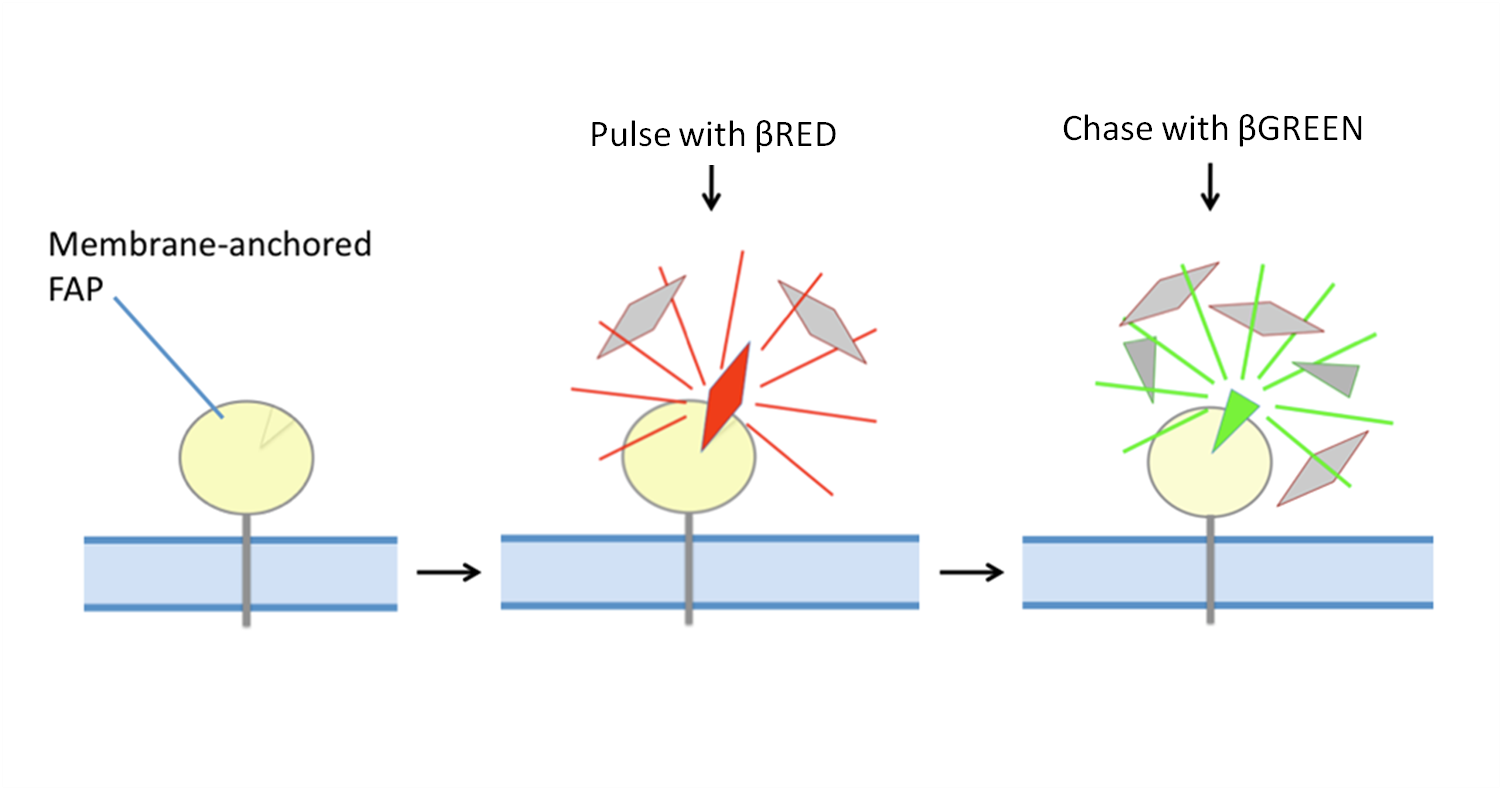
The basic experiment can be illustrated in this way:
Both fluorogens in this case are impermeable and the βGREEN fluorogen will quickly replace the βRED giving us two populations of the same protein. The “Red” population represents protein that was at the surface when the Red fluorogen was applied, internalized and has not yet returned to the surface. The “Green” population will be from protein that stayed at the surface or returned to the surface after the addition of green fluorogen.
It then becomes very straightforward to perform internalization assays utilizing a cell line line expressing a FAP-tagged GPCR.
1. Add test compound plus red fluorogen
2. Allow time for response
3. Add green fluorogen
4. Read
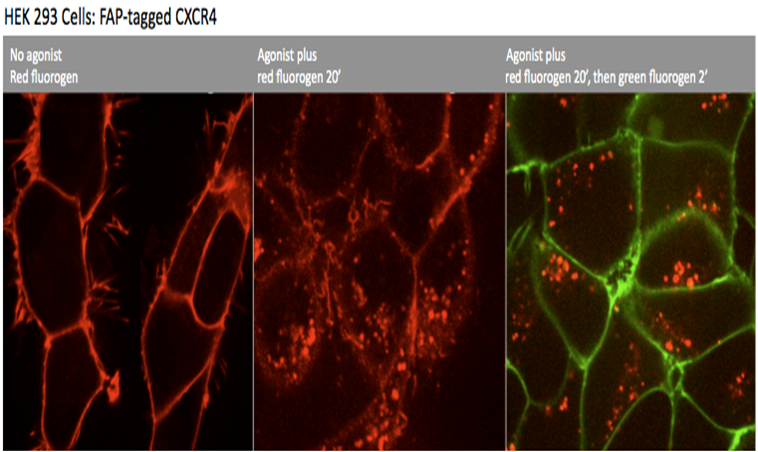
An example is shown in the images below:
The left panel shows CXCR4 at the surface before the addition of agonist. The middle panel shows the image 20 minutes after the addition of agonist, clearly showing a significant response. The final panel shows the image after the addition of the chase fluorogen (green). The red color in the final image represents internalized protein and the green represents protein that remained at the surface.
This data can easily be quantified. Below shows a dose response and a time response curves for CXCR4 when stimulated with CXCL12.
Antagonist, Selectivity, and Re-sensitization Assays can easily be run utilizing the same experimental design.
SpectraGenetics has tagged more than 150 GPCRs and validated more than 30 cell lines.